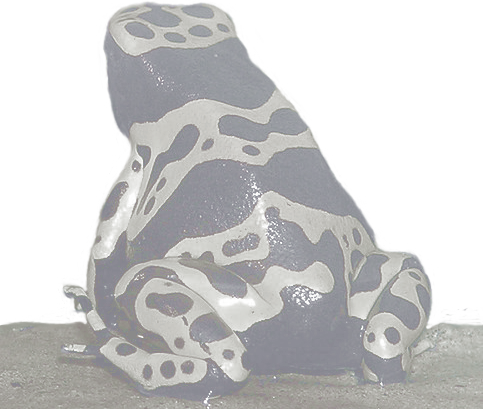
This post will provide a general introduction to the biology of predator-prey systems: I will start with the evolution of predator and prey and their consequential co-evolution in predator-prey systems. The focus will then move to mechanisms to avoid attack and how it results in a complex signalling system called aposematism. I will present the scientific background to the emergence of aposematism and lay out the open questions which motivate the following chapters.
 The application of temporal difference learning
The application of temporal difference learningAn experience-based aversive learning model of foraging behaviour in uncertain environments is presented. We use Q-learning as a model-free implementation of Temporal difference learning motivated by growing evidence for neural correlates in natural reinforcement settings.
 MicroRNA seed sites
MicroRNA seed sitesAn analysis of the structural context of microRNA seed sites within 3’UTR sequences points to certain microRNA families which show enriched seed sites within hairpin structures. A sub-set of these hairpins are enriched by AU rich PUM2 binding motifs.
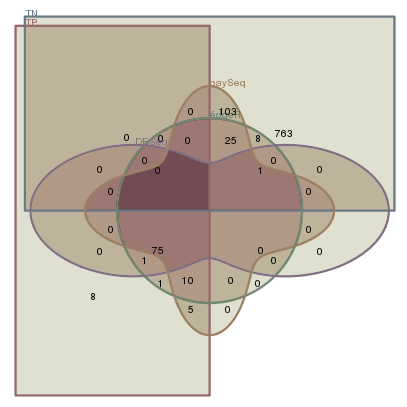
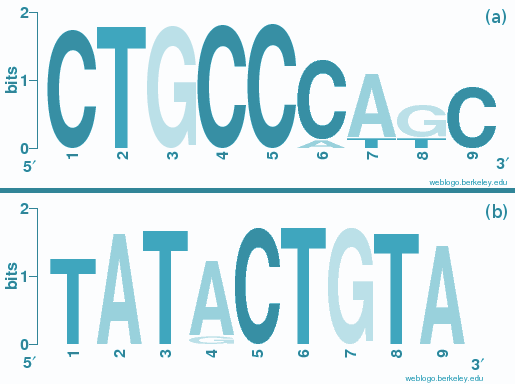 Analysing Deep Sequencing Data
Analysing Deep Sequencing DataI used three methods on simulated data-sets to evaluate their performance. The simplest approach is by the edgeR method which assumes a linear relation of mean and dispersion. The DESeq method extends this approach to an non-linear data-driven dependency. The baySeq method uses general Bayesian methods to estimate posterior probabilities.
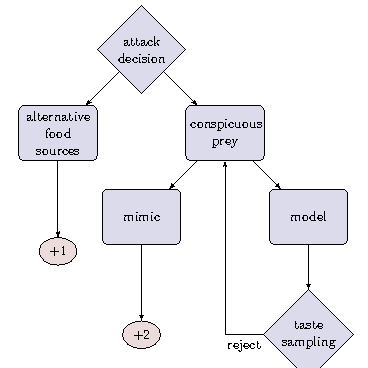
 The Evolutionary Dynamics of Aposematism
The Evolutionary Dynamics of AposematismThe majority of species are under predatory risk in their natural habitat and targeted by predators as part of the food web. During the evolution of ecosystems, manifold mechanisms have emerged to avoid predation.
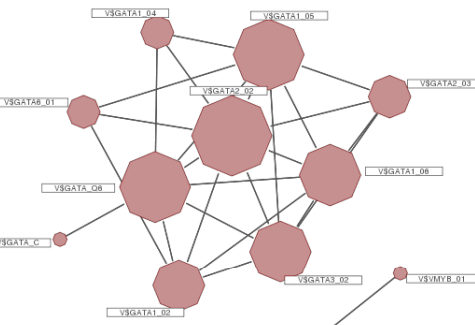
 Bioinformatic Prediction of Transcription Factor Binding Sites within short DNA Sequences
Bioinformatic Prediction of Transcription Factor Binding Sites within short DNA SequencesSince fast and cheap DNA sequencing methods have become firmly established in a wide range of biology the identification of functional elements and transcription factor binding sites (TFBSs) remains an immense challenge.